See on Scoop.it – Amazing Science
Scientists have obtained the first detailed molecular structure of a member of the Tet family of enzymes.
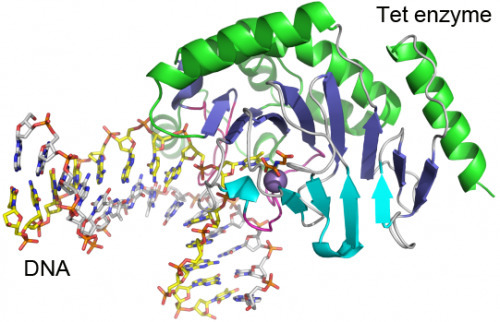
Cytosine residues in mammalian DNA occur in five forms: cytosine (C), 5-methylcytosine (5mC), 5-hydroxymethylcytosine (5hmC), 5-formylcytosine (5fC) and 5-carboxylcytosine (5caC). The ten-eleven translocation (Tet) dioxygenases convert 5mC to 5hmC, 5fC and 5caC in three consecutive, Fe(II)- and α-ketoglutarate-dependent oxidation reactions [1, 2, 3, 4]. The Tet family of dioxygenases is widely distributed across the tree of life [5], including in the heterolobosean amoeboflagellate Naegleria gruberi. The genome of Naegleria [6] encodes homologues of mammalian DNA methyltransferase and Tet proteins [7]. Here we study biochemically and structurally one of the Naegleria Tet-like proteins (NgTet1), which shares significant sequence conservation (approximately 14% identity or 39% similarity) with mammalian Tet1. Like mammalian Tet proteins, NgTet1 acts on 5mC and generates 5hmC, 5fC and 5caC. The crystal structure of NgTet1 in complex with DNA containing a 5mCpG site revealed that NgTet1 uses a base-flipping mechanism to access 5mC. The DNA is contacted from the minor groove and bent towards the major groove. The flipped 5mC is positioned in the active-site pocket with planar stacking contacts, Watson–Crick polar hydrogen bonds and van der Waals interactions specific for 5mC. The sequence conservation between NgTet1 and mammalian Tet1, including residues involved in structural integrity and functional significance, suggests structural conservation across phyla.
The finding is important for the field of epigenetics because Tet enzymes chemically modify DNA, changing signposts that tell the cell’s machinery “this gene is shut off” into other signs that say “ready for a change.”
Tet enzymes’ roles have come to light only in the last five years; they are needed for stem cells to maintain their multipotent state, and are involved in early embryonic and brain development and in cancer.
Researchers led by Xiaodong Cheng, PhD, determined the structure of a Tet family member from Naegleria gruberi by X-ray crystallography. The structure shows how the enzyme interacts with its target DNA, bending the double helix and flipping out the base that is to be modified. “This base flipping mechanism is also used by other enzymes that modify and repair DNA, but we can see from the structure that the Tet family enzymes interact with the DNA in a distinct way,” Cheng says. Mammalian Tet enzymes appear to have an additional regulatory domain that the Naegleria forms do not; understanding how that domain works will be a new puzzle opened up by having the Naegleria structure, Cheng says.
See on phys.org